Infrared-optical hybrid “sees” molecular signatures of cancer biopsies
NIBIB-funded bioengineers at the University of Illinois have combined standard microscopy, infrared light, and artificial intelligence (AI) to assemble digital biopsies that identify important molecular characteristics of cancer biopsy samples without dyes or labels.
The standard for cancer biopsies is treating the tissue to be tested with staining solutions that reveal cellular details that help a pathologist identify whether a sample is cancerous, and if so, give some idea of cancer severity. Fixing and staining the tissue can take several hours, and although pathologists are very skilled, making a diagnosis based on the size and shape of cells—enhanced by the staining process—can differ between individuals.
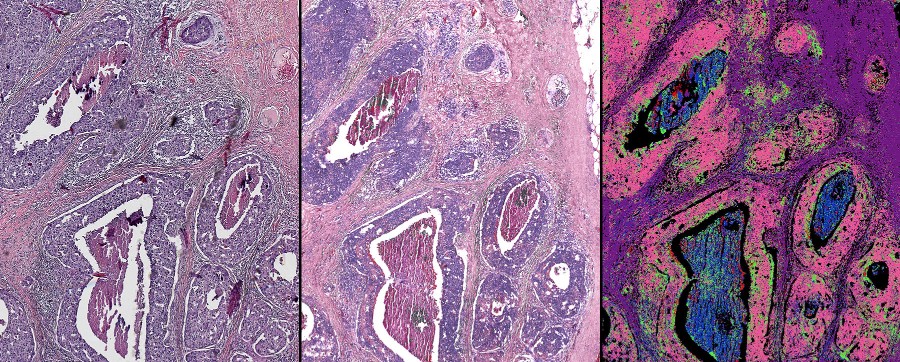
In an effort to obtain more rapid, comprehensive, and objective information from biopsy samples, bioengineers at the University of Illinois have built a unique microscope that combines standard visible light microscopy with infrared light. The research is led by Rohit Bhargava, Ph.D., a professor of bioengineering and the founding director of the Cancer Center at Illinois. The team is using machine learning, a type of AI, to process the signals from the hybrid microscope to create unique images of cancer samples that carry much more information than a standard pathology stain.
The technique was created by adding an infrared laser and specialized lenses to the standard light microscope found in labs and clinics. The set-up captures both high resolution optical images and infrared molecular signals emitting from a sample bathed in white and infrared light.
“We built the hybrid microscope with commercially available components,” explained Bhargava. “Numerous labs with optical microscopes will be able to build similar instruments, which will help this new approach to be disseminated widely and rapidly.”
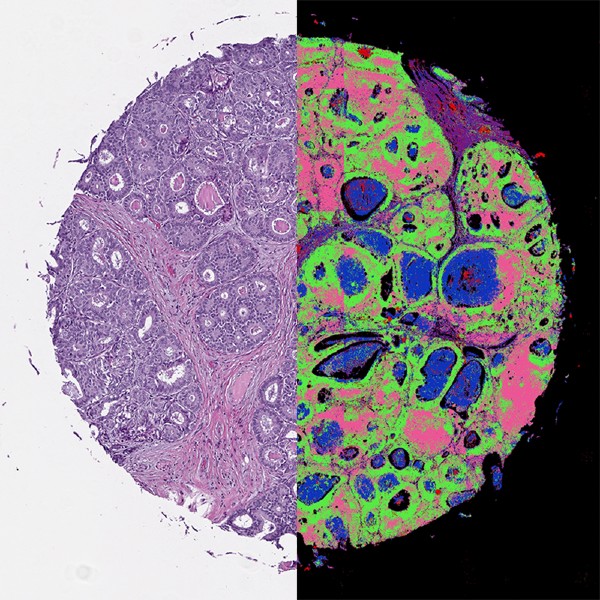
A key component of the new system is a machine learning program. The computer program combines the white light and infrared signals that bounce off the biopsy sample under the microscope. The result is an image that mimics the type obtained if the sample were stained in a traditional pathology lab—created in minutes instead of hours.
The pathology dye stain and the digital “stain” constructed by AI both highlight the size and pattern of the cells in the tissue, with certain patterns being characteristic of cancerous tissues. However, AI gathers information contained in the sample that cannot be seen with the human eye.
“Cancer cells have differences in their chemical composition and metabolism”, explained Bhargava. “Information from the infrared signal gives a readout of the molecular composition of the tissues that can be analyzed by AI for decision-making in pathology.”
The extra molecular data reveals specific characteristics of the tumor and the surrounding healthy tissues that can provide better and more consistent information about the cancer’s progression.
Because infrared light has multiple wavelengths with different chemical information, the researchers are refining the AI computational programs so it can analyze the different types of cells and diseases. The goal is to achieve extremely precise and reproducible “cancer mapping.”
The technique promises to speed up results, reduce costs and provide what Bhargava calls an “all-digital” analysis of cancer pathology. The team is also exploring use of the hybrid microscope for additional biomedical applications such as forensics and polymer science.
The work was reported in the Proceedings of the National Academy of Sciences1. Funding was provided by the National Institute of Biomedical Imaging and Bioengineering through grants R01EB009745 and T32EB019944. Additional funding was provided through an Agilent Thought Leader award, a Marie Curie Fellowship and a Beckman Institute graduate fellowship.
