- What is computational modeling?
- How can computational modeling improve medical care and research?
- What are digital twins?
- How are NIBIB-funded researchers using computational modeling to improve health?
What is computational modeling?
Computational modeling is the use of computers to simulate and study complex systems using mathematics, physics, and computer science.
Complex systems are characterized by numerous variables (factors) that can affect how the system functions, which can ultimately influence its outcomes. Using computational modeling, complex systems are studied in a virtual environment using variables that define each system. The computer model then simulates the system under different conditions, creating simulated outputs or predictions. For example, weather forecasting models make predictions based on numerous atmospheric variables, and flight simulators use complex equations that govern how aircraft fly and react to variables such as turbulence, air density, and precipitation.
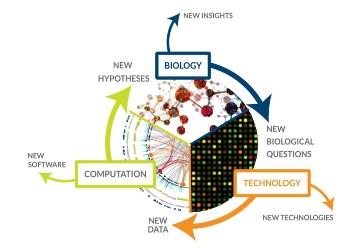
In biomedical research, computer modeling allows scientists to conduct thousands of simulated experiments by computer. The thousands of computer experiments can identify the handful of laboratory experiments that are most likely to improve scientific understanding, saving both time and money.
There are two broad approaches to modeling real-world systems: mechanistic modeling and data-driven modeling. Mechanistic models are based on an underlying understanding of how a system works and are built using established scientific principles (such as the laws of physics and known biochemical processes). Data-driven models, on the other hand, leverage patterns and associations observed in vast datasets to predict how complex systems operate without explicit knowledge of how they work. Many computational models use both of these approaches and are known as hybrid models.
Today’s computational models can study a biological system at multiple levels, ranging from molecules to tissues to entire organisms. Models of how disease develops include molecular processes, cell-to-cell interactions, and how these changes affect tissues and organs. Studying systems at multiple levels is known as multiscale modeling.
How can computational modeling improve medical care and research?
Clinical decision support. Computational models intelligently gather, filter, analyze, and present health information to provide guidance to doctors for disease treatment based on detailed characteristics of each patient. The systems help to provide informed and consistent care of a patient as they transfer to appropriate hospital facilities and departments and receive various tests during their course of treatment.
Predicting drug side effects. Researchers use computational modeling to help design drugs that will be the safest for patients and least likely to have side effects. The approach can augment the use of animal models and potentially reduce the many years needed to develop safe and effective medications.
Tracking infectious diseases. Computational models are being used to track infectious diseases in populations, identify the most effective interventions, and monitor and adjust interventions to reduce the spread of disease. Identifying and implementing interventions that curb the spread of disease are critical for saving lives and reducing stress on the health care system during infectious disease outbreaks.
What are digital twins?
Digital twins are an emerging technology that pair computational models with a physical counterpart to develop an evolving and dynamic framework that is continuously updated to enable predictions and inform decisions about a complex system. According to the National Academies of Sciences, Engineering, and Medicine (NASEM), digital twin technologies consist of a real-life (physical) representation of a system that is “twinned” with a virtual representation of that system. These representations are linked with bidirectional information exchange to provide optimal decision support.
One potential application for digital twin technology includes a patient with a disease, such as cancer. Here’s how it would work: A digital twin is built using clinical assessments from the real-world patient, such as lab tests, tissue specimens, and medical images. As the patient’s disease evolves—for example, as a tumor grows or shrinks—the associated medical information from the physical world is used to continuously update the digital twin. This digital twin can be used to explore a host of different treatment options by simulating a variety of therapeutic regimens and predicting clinical outcomes. Continuous communication between the physical and digital components throughout the course of disease could facilitate the real-time adjustment of a personalized treatment plan with the highest likelihood of success.
While computational models are an important aspect of digital twin methodology, they are only one component of this emerging technology.
How are NIBIB-funded researchers using computational modeling to improve health?
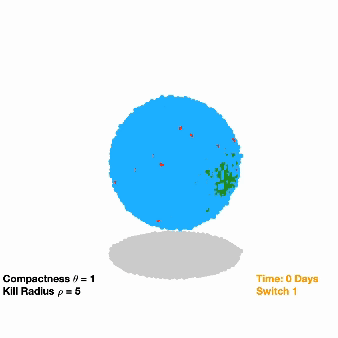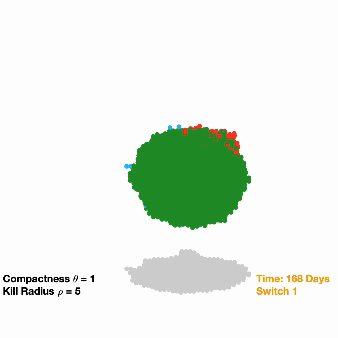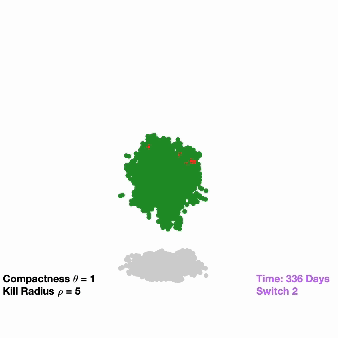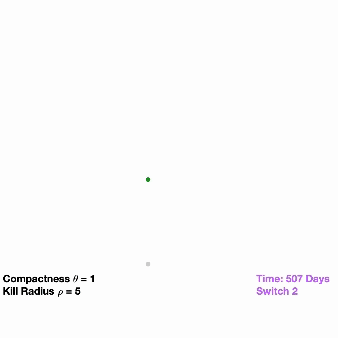
Switching off drug-resistant cancer: Due to their eclectic mix of mutations, cancer cells have the ability to outwit therapeutic drugs, developing resistance to certain treatments and leaving behind durable cell populations. To take advantage of this evolutionary trajectory, researchers are investigating a treatment that involves inserting a pair of synthetic genes into a small pool of cancer cells. The first gene confers resistance to a specific anti-cancer drug, while the second gene causes these resistant cells (and those around them) to die once they’ve become the dominant population in the tumor. Researchers used computational models to understand the feasibility of this approach and also to fine-tune their strategy for a specific cancer type before moving their work into cancer cells and eventually animal models.
Modeling fluid dynamics in the human heart: Cardiovascular disease is the leading cause of death in the United States and around the world. Models of the heart are essential preclinical tools to better understand both normal and irregular cardiac function and to ultimately design and test lifesaving interventions. While comprehensive computational models of the heart exist, none currently include three-dimensional representations of the four cardiac valves, which are the one-way gates that direct blood flow through the chambers of the heart. Cardiac valve performance can decrease as we age and is linked with a host of cardiovascular diseases, underscoring the need for their accurate representation in computational models. In this study, the researchers introduce a fluid-structure interaction model that provides detailed representations of all four cardiac valves. Their tool can effectively model fluid dynamics of the human heart that are in excellent agreement with clinical and experimental data. Detailed cardiac computational models like these can provide novel ways to study cardiac dysfunction, such as congenital defects and heart failure, which are challenging or impossible to perform in human patients.
A multiscale model of the primary motor cortex: The brain is a highly complex organ, and understanding its abilities requires studying the interaction of its various components at multiple scales, ranging from the molecular to the behavioral. The locations and connections of different neurons in the brain affect cortical function, yet detailed multiscale models of the primary motor cortex, the region of the brain responsible for voluntary movement, are scarce. In this study, researchers report the development of a multiscale model of mouse primary motor cortex with over 10,000 neurons and 30 million synapses. This model incorporates physiological and anatomical data and can faithfully predict mouse neural responses associated with behavioral states (such as quiet wakefulness and voluntary movement), setting the stage for future synthetic experiments. Their model is freely available as a research tool and can be updated and expanded with new data, such as those collected from the NIH BRAIN Initiative® Cell Census Network.
Tools for personalized treatments for neuromuscular conditions: It is estimated that neuromuscular injuries affect nearly 20% of the U.S. adult population. Existing standardized interventions are limited, as each injury requires unique treatment needs. NIH-funded researchers have created a freely available user software system called OpenSim that facilitates the modeling of musculoskeletal structures, which can be used to learn more about biomechanics and develop treatment plans. The researchers recently reported two new toolsets that add model personalization and treatment optimization functionalities to their existing platform. Using data collected from a post-stroke individual with impaired walking function, the researchers used this pipeline to develop a personalized musculoskeletal model and identified how the individual’s neural control could be adapted to improve their gait. Future work could include predictive simulations to help design optimized, personalized treatments for patients with neuromuscular conditions.
Updated September 2025
