New system can model four tissues on one chip with one overarching goal—more accurate models of human physiology
Tissue chips—tiny mimics of human organs, just millimeters in size—represent an alternative to animal models as a way to study disease or evaluate drugs. However, a major limitation of tissue chips is that they do not faithfully imitate tissue interactions, so it’s impossible to know how a treatment for liver disease, for example, might affect another organ, like the heart.
To improve this technology, NIBIB-funded researchers have developed an interlinked tissue chip system that can model four mature organs in their perspective environments simultaneously. These multi-organ tissue chips, which could be personalized to model individual patients, may represent a new way to evaluate the systemic effects of novel drugs.
“As a field, we are constantly aiming to develop true-to-life models that are accurate representations of human physiology,” said David Rampulla, Ph.D., director of the division of Discovery Science & Technology at NIBIB. “This study outlines a new method to model interconnected human organs, which could allow researchers to better investigate diseases—or treatments—that affect multiple different tissues.”
Tissue chip systems that simultaneously model multiple organs have been previously explored, but here’s the catch—the different tissues were all grown in the same environment, which is not representative of the human body. Each of our organs has unique environmental requirements to function optimally—different amounts of salts, acids, proteins, growth factors, and hormones, for example. A collection of organ models that are all grown in the same conditions in the lab can have altered phenotypes and fail to accurately represent mature human tissues.
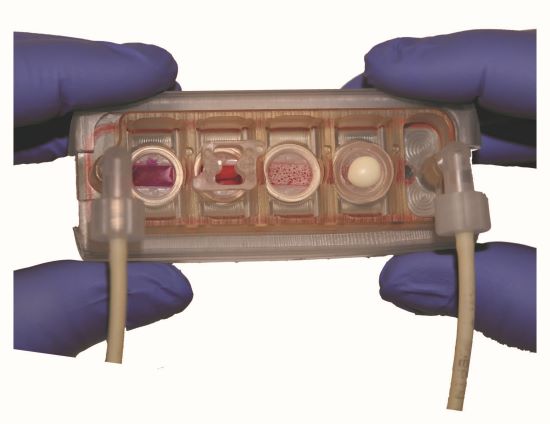
A team of bioengineers from Columbia University has bridged the gap between growing tissues in isolation and growing tissues in a shared environment. They have developed an interlinked system where each organ is grown in a separate chamber, but all the chambers are connected by vascular flow (similar to organs connected by blood circulating through the body). What’s more, the chambers are separated by a semi-permeable, vascular barrier, allowing the separate tissues to grow in their optimal environment, but also allowing for communication between the different organs (such as the exchange of cells, immune signals, and secreted factors).
“Our goal was to develop an improved model of human physiology, so we took inspiration from the human body, which relies on inter-organ crosstalk while maintaining ideal tissue environments,” explained first study author Kacey Ronaldson-Bouchard, Ph.D., an associate research scientist at Columbia University in New York. “By developing a multi-organ platform where tissues can ‘talk’ to one another through secreted factors, we can more accurately model systemic conditions and disease progression.”
In their study, published in Nature Biomedical Engineering, the research team reported on a multi-organ chip that contained four matured human tissues: heart, liver, bone, and skin, all linked through vasculature to form an interconnected system. After the matured organs grew for four weeks in the interlinked chip, all maintained their proper phenotypes, providing faithful models of human tissue. The researchers also demonstrated communication between the different organs: when they injured the heart tissue, they found cardiac-specific markers in the other three tissues. Additionally, when they added immune cells into the vascular flow, they found that these cells migrated to the damaged heart tissue, but not to the healthy organs.
To evaluate the ability of their system to model drug testing and toxicity, the researchers studied doxorubicin, a common chemotherapeutic used in cancer treatments. They found that the drug behaved similarly in their tissue chip system as it does in human patients, including the side effect of cardiotoxicity. They believe that additional studies with their system could lead to an improved method to evaluate novel therapies.
Ronaldson-Bouchard noted that their platform is amenable to any engineered tissue, not just the four that were evaluated here. “Scientists can simply ‘plug in’ tissues of interest—either healthy or diseased human tissues—to support multiple stages of preclinical development,” she said. Further, by using patient-derived cells to grow matured tissues, their system could tease out how a specific disease might affect patient populations differently (based on ethnicity, age, sex, or other factors). This type of information could improve drug efficacy and better inform clinical trial design, she said.
“Systemic conditions—such as cancer metastasis, multi-organ drug toxicity, infection, and aging—are largely left out of the drug development pipeline and lack adequate treatments, primarily due to the lack of accurate models,” said senior study author Gordana Vunjak-Novakovic, Ph.D., University Professor and Mikati Foundation Professor of Biomedical Engineering and Medical Sciences at Columbia University. “There is a need for more relevant models, particularly ones that facilitate understanding of how a drug will work in specific patient populations. Our multi-organ chip demonstrates the potential to do just this.”
This study was supported by grants from NIBIB (UG3EB025765 and P41EB027062), the National Cancer Institute (NCI; grants R01CA249799, R35CA197745, and P30CA013696), the National Center for Advancing Translational Sciences (NCATS; grant UL1TR001873), and the NIH Office of the Director (OD; grants (S10OD012351 and S10OD021764).
Study reference: Ronaldson-Bouchard K, Teles D, Yeager K, et al. A multi-organ chip with matured tissue niches linked by vascular flow. Nat Biomed Eng. 2022;6(4):351-371. doi:10.1038/s41551-022-00882-6
